IBEA¶
Example¶
[1]:
from jmetal.algorithm.multiobjective.ibea import IBEA
from jmetal.operator.crossover import SBXCrossover
from jmetal.operator.mutation import PolynomialMutation
from jmetal.problem import ZDT1
from jmetal.util.termination_criterion import StoppingByEvaluations
problem = ZDT1()
max_evaluations = 25000
algorithm = IBEA(
problem=problem,
kappa=1.,
population_size=100,
offspring_population_size=100,
mutation=PolynomialMutation(probability=1.0 / problem.number_of_variables, distribution_index=20),
crossover=SBXCrossover(probability=1.0, distribution_index=20),
termination_criterion=StoppingByEvaluations(max_evaluations)
)
algorithm.run()
front = algorithm.get_result()
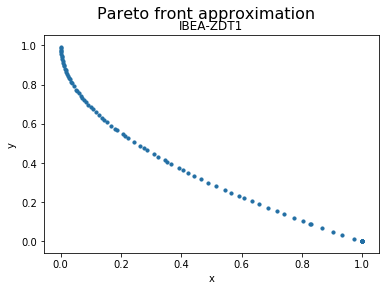
We can now visualize the Pareto front approximation:
[3]:
from jmetal.lab.visualization.plotting import Plot
plot_front = Plot(plot_title='Pareto front approximation', axis_labels=['x', 'y'])
plot_front.plot(front, label='IBEA-ZDT1')
API¶
- class jmetal.algorithm.multiobjective.ibea.IBEA(problem: ~jmetal.core.problem.Problem, population_size: int, offspring_population_size: int, mutation: ~jmetal.core.operator.Mutation, crossover: ~jmetal.core.operator.Crossover, kappa: float, termination_criterion: ~jmetal.util.termination_criterion.TerminationCriterion = <jmetal.util.termination_criterion.StoppingByEvaluations object>, population_generator: ~jmetal.util.generator.Generator = <jmetal.util.generator.RandomGenerator object>, population_evaluator: ~jmetal.util.evaluator.Evaluator = <jmetal.util.evaluator.SequentialEvaluator object>)[source]¶
Bases:
GeneticAlgorithm[S,R]- create_initial_solutions() List[S][source]¶
Creates the initial list of solutions of a metaheuristic.
